Beischlag Lab
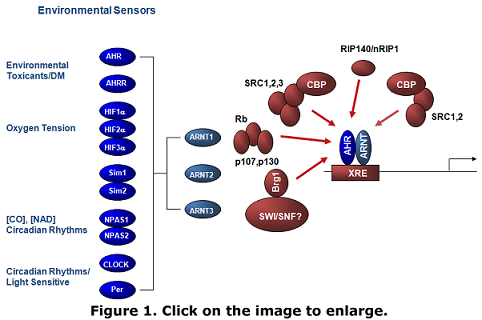
All living organisms share the requirement for tightly regulated gene expression and the ability to respond to external signals to ensure the precise spatial and temporal control of gene activation. Transcription factors are proteins that are essential for the activation or repression of gene expression. The basic-Helix-Loop-Helix (bHLH)-PAS domain super-family of transcription factors is composed of a group of structurally related proteins that are thought to serve as sensors of environmental cues (Figure 1). Proteins in this family are responsive to environmental toxicants (aryl hydrocarbon receptor, AHR), and oxygen tension (hypoxia-inducible factors, HIF-a/ß). My lab studies two distinct aspects of transcription factor function; (i) the molecular mechanisms of transcription factor tethering, or cross-talk, between the nuclear receptor transcription factor, estrogen receptor-a(ERa), and AHR, and; (ii) the regulation of co-activator function during hypoxia-inducible transcription. |
 |
These systems offer several advantages. First, because they are structurally similar, corollaries of function are easier for comparison as opposed to observations made between structurally dissimilar classes of transcription factors. Second, they share structural similarities with a wider family of bHLH transcription factors, and functional similarities with other ligand activated transcription factors, in particular, nuclear receptors. Thus, mechanistic observations of bHLH-PAS protein function will increase our understanding transcription factor function, in general. Finally, activation of both factors leads to the robust and tightly regulated activation of their respective prototypical target genes, VEGF and EPO for HIF-1a, and CYP1A1 for AHR. In the absence of HIF-1a and AHR activation, there is virtually no measurable transcriptional activity of these genes, but upon activation there is a robust increase in messenger (mRNA) levels. This easily identifiable and quantifiable readout greatly facilitates the study of phenomena that would tend to repress or attenuate transcription factor function. The long-term objective of my research program is to understand how proteins recruited to, or displaced from promoters during activated transcription mediate the appropriate spatial and temporal control of gene expression.
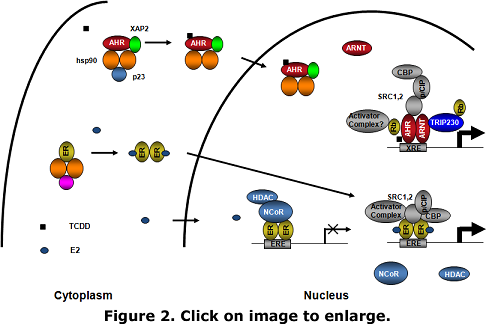
Normal estrogen physiology depends on the appropriate spatial and temporal activation of target genes regulated by estrogen receptors (ER's). The inappropriate expression of ER target genes leads to uncontrolled cell proliferation in certain tissues such as the breast, promoting tumor growth. To activate gene transcription, ER must assemble specific proteins including other transcription factors, such as the aryl hydrocarbon receptor (AHR), to the regulatory regions of target genes (Figure 2). Despite mounting evidence indicating that activation of AHR inhibits estrogen-dependent tumor growth, little is known regarding the molecular mechanisms that regulate this phenomenon. Elucidating the mechanisms by which AHR modulates ER function is important for understanding how transcription factor cross-talk affects normal and deregulated gene expression and cell growth. Understanding these mechanisms and the gene expression programs involved offers the potential for novel therapies based on the manipulation of specific factors or gene expression pathways. It is our goal to define the role of AHR in the etiology of ER-positive (ER+) breast cancers, and to explore the potential therapeutic value of certain AHR ligands. We are pursing avenues of research to elucidate the molecular determinants of ER-AHR cross-talk and the gene expression programs affected by this interaction. |
 |
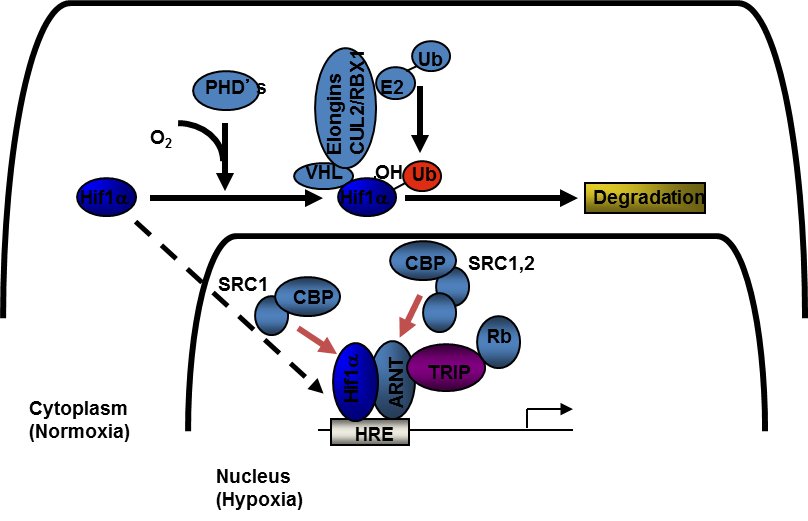
All cells respond to hypoxia. Perhaps the best characterized hypoxic response regulated by HIF-1α/ARNT(HIF-1β) is the formation new vasculature, or angiogenesis. The blood carries oxygen to all tissues and the only means by which growing tissues can improve their oxygen supply is by the de novo formation of vascular beds. This is particularly important for tumor biology. As solid tumors grow, their requirement for oxygen surpasses the available supply; as such, solid tumors have regions that are chronically hypoxic. This leads to the activation/up-regulation of HIF-1α, and it activates transcription of hypoxia responsive target genes that initiate a wide variety of cellular and physiological programs including angiogenesis. It has been shown that HIF-1α-regulated is associated with tumor growth and metastasis. We are interested in the molecular determinants underlying HIF-1α mediated gene transcription. How HIF-1α is regulated during normoxia or normal oxygen tension is fairly well understood. However, little is known of the factors involved in HIF-1α-regulated transcription during hypoxia, or periods of low oxygen tension (Figure 3). We are currently investigating the role of a transcriptional co-activator, TRIP230 among other proteins, in HIF-1α regulated processes. In addition, genes batteries up or down-regulated in response to hypoxia in different tumor types are still largely unknown. We are actively pursuing these avenues of research. |
 |