Chapter Contents
Previous
Next
|
Chapter Contents |
Previous |
Next |
| The REG Procedure |
This section discusses the INFLUENCE option, which produces several influence statistics, and the PARTIAL option, which produces partial regression leverage plots.
Belsley, Kuh, and Welsch propose a cutoff of 2p/n, where n is the number of observations used to fit the model and p is the number of parameters in the model. Observations with hi values above this cutoff should be investigated.
For each observation, PROC REG first displays the residual, the studentized residual (RSTUDENT), and the hi. The studentized residual RSTUDENT differs slightly from STUDENT since the error variance is estimated by s(i)2 without the ith observation, not by s2. For example,
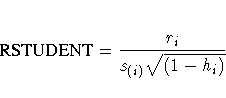
Observations with RSTUDENT larger than 2 in absolute value may need some attention.
The COVRATIO statistic measures the change in the determinant of the covariance matrix of the estimates by deleting the ith observation:
Belsley, Kuh, and Welsch suggest that observations with

where p is the number of parameters in the model and n is the number of observations used to fit the model, are worth investigation.
The DFFITS statistic is a scaled measure of the change in the predicted value for the ith observation and is calculated by deleting the ith observation. A large value indicates that the observation is very influential in its neighborhood of the X space.
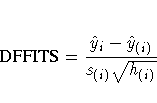
Large values of DFFITS indicate influential observations.
A general cutoff to consider is 2; a size-adjusted
cutoff recommended by Belsley, Kuh, and Welsch is
![]() , where n and p are as defined previously.
, where n and p are as defined previously.
The DFFITS statistic is very similar to Cook's D, defined in the section "Predicted and Residual Values".
The DFBETAS statistics are the scaled measures of the change in each parameter estimate and are calculated by deleting the ith observation:
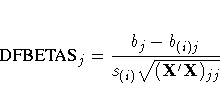
where (X'X)jj is the (j,j)th element of (X'X)-1.
In general, large values of DFBETAS indicate observations
that are influential in estimating a given parameter.
Belsley, Kuh, and Welsch recommend 2 as a general
cutoff value to indicate influential observations
and ![]() as a size-adjusted cutoff.
as a size-adjusted cutoff.
Figure 55.42 shows the tables produced by the INFLUENCE option for the population example (the section "Polynomial Regression"). See Figure 55.29 for the fitted regression equation.
proc reg data=USPopulation;
model Population=Year YearSq / influence;
run;
In Figure 55.42, observations 16, 17, and 19 exceed the cutoff value of 2 for RSTUDENT. None of the observations exceeds the general cutoff of 2 for DFFITS or the DFBETAS, but observations 16, 17, and 19 exceed at least one of the size-adjusted cutoffs for these statistics. Observations 1 and 19 exceed the cutoff for the hat diagonals, and observations 1, 2, 16, 17, and 18 exceed the cutoffs for COVRATIO. Taken together, these statistics indicate that you should look first at observations 16, 17, and 19 and then perhaps investigate the other observations that exceeded a cutoff.
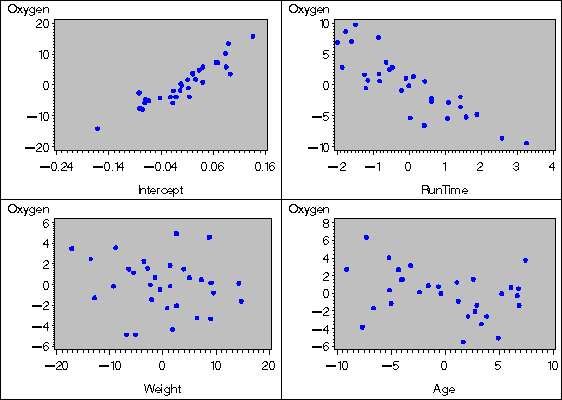
For a given regressor, the partial regression leverage plot is the plot of the dependent variable and the regressor after they have been made orthogonal to the other regressors in the model. These can be obtained by plotting the residuals for the dependent variable against the residuals for the selected regressor, where the residuals for the dependent variable are calculated with the selected regressor omitted, and the residuals for the selected regressor are calculated from a model where the selected regressor is regressed on the remaining regressors. A line fit to the points has a slope equal to the parameter estimate in the full model.
In the plot, points are marked by the number of replicates appearing at one position. The symbol '*' is used if there are ten or more replicates. If an ID statement is specified, the left-most nonblank character in the value of the ID variable is used as the plotting symbol.
The following statements use the fitness data in Example 55.1 with the PARTIAL option to produce the partial regression leverage plots in the OUTPUT window. The plots are not shown.
proc reg data=fitness lineprinter;
model Oxygen=RunTime Weight Age / partial;
run;
The following statements create one of the partial regression
plots on a high resolution graphics device for the fitness data;
all four plots (created by regressing Oxygen and one of the
variables on the remaining variables) are displayed
in Figure 55.43.
Notice that the Int variable is explicitly added to be
used as the intercept term.
data fitness2;
set fitness;
Int=1;
proc reg data=fitness2 noprint;
model Oxygen Int = RunTime Weight Age / noint;
output out=temp r=ry rx;
symbol1 c=blue;
proc gplot data=temp;
plot ry*rx / cframe=ligr;
label ry='Oxygen'
rx='Intercept';
run;
|
|
Chapter Contents |
Previous |
Next |
Top |
Copyright © 1999 by SAS Institute Inc., Cary, NC, USA. All rights reserved.