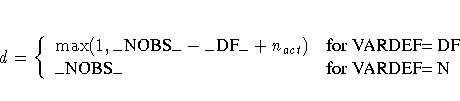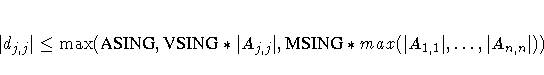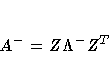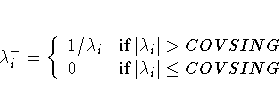Covariance Matrix
The COV= option must be specified to compute an approximate
covariance matrix for the parameter estimates under asymptotic
theory for least-squares, maximum-likelihood, or Bayesian
estimation, with or without corrections for degrees of freedom
as specified by the VARDEF= option.
Two groups of six different forms of covariance matrices
(and therefore approximate standard errors) can be computed
corresponding to the following two situations:
- The LSQ statement is specified, which means that
least-squares estimates are being computed,
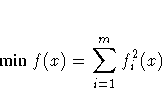
- The MIN or MAX statement is specified, which means
that maximum-likelihood or Bayesian estimates are
being computed,
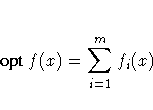
where opt is either min or max.
In either case, the following matrices are used:
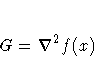
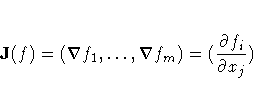
-
JJ(f) = J(f)T J(f)
-
V = J(f)T Diag(fi2) J(f)
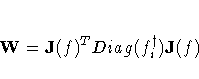
where
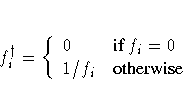
For unconstrained minimization, or when none of the final
parameter estimates is subjected to linear equality or active
inequality constraints, the formulas of the six types of
covariance matrices areas follows.
|
COV
|
MIN or MAX Statement
|
LSQ Statement
|
| 1 | M | [ _NOBS_/d] G-1 JJ(f) G-1 | [ _NOBS_/d] G-1 V G-1 |
| 2 | H | [ _NOBS_/d] G-1 | 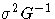 |
| 3 | J | [1/d] W-1 | 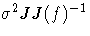 |
| 4 | B | [1/d] G-1 W G-1 | 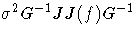 |
| 5 | E | [ _NOBS_/d] JJ(f)-1 | [1/d] V-1 |
| 6 | U | [ _NOBS_/d] W-1 JJ(f) W-1 | [ _NOBS_/d] JJ(f)-1 V JJ(f)-1 |
The value of d depends on the VARDEF= option and on the value
of the _NOBS_ variable:
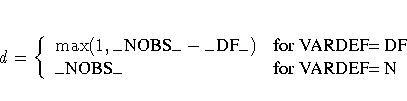
where _DF_ is either set in the program statements or set
by default to n (the number of parameters) and _NOBS_ is
either set in the program statements or set by default to
nobs * mfun; nobs is
the number of observations in the data
set and mfun is the number of functions listed in the LSQ,
MIN, or MAX statement.
The value 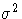 depends on the specifications of the SIGSQ=
options and on the value of d:
depends on the specifications of the SIGSQ=
options and on the value of d:
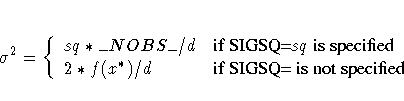
where f(x*) is the value of the objective function at the
optimal parameter estimates x*.
The two groups of formulas distinguish between two situations:
- For least-squares estimates, the error
variance can be estimated from the objective function value and
is used in three of the six different forms of covariance
matrices. If you have an independent
estimate of the error variance, you can specify it with the
SIGSQ= option.
- For maximum-likelihood or Bayesian estimates,
the objective function should be the logarithm of the likelihood
or of the posterior density when using the MAX statement.
For minimization, the inversion of the matrices in these formulas
is done so that negative eigenvalues are considered zero,
resulting always in a positive semidefinite covariance matrix.
In small samples, estimates of the covariance matrix based on
asymptotic theory are often too small and should be used with
caution.
If the final parameter estimates are subjected to nact > 0
linear equality or active linear inequality constraints, the
formulas of the covariance matrices are modified
similar to Gallant (1987) and Cramer (1986, p. 38) and
additionally generalized for applications with singular
matrices. In the constrained case, the value of d used in
the scalar factor 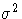 is defined by
is defined by
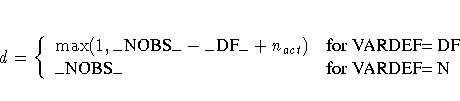
where nact is the number of active constraints, and
_NOBS_ is set as in the unconstrained case.
For minimization, the covariance matrix should be positive
definite; for maximization it should be negative definite.
There are several options available to check for a rank
deficiency of the covariance matrix:
- The ASINGULAR=, MSINGULAR=, and VSINGULAR= options can
be used to set three singularity criteria for the inversion
of the matrix A needed to compute the covariance matrix,
when A is either the Hessian
or one of the crossproduct Jacobian matrices.
The singularity criterion used for the inversion is
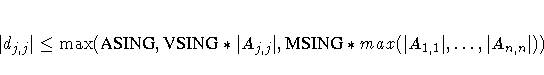
where dj,j is the diagonal pivot of the matrix A,
and ASING, VSING and MSING are the specified values of
the ASINGULAR=, VSINGULAR=, and MSINGULAR= options. The
default values are
- ASING: the square root of the smallest positive double
precision value
- MSING: 1e-12 if the SINGULAR= option is not specified
and
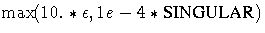 otherwise,
where
otherwise,
where 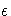 is the machine precision
is the machine precision
- VSING: 1e-8 if the SINGULAR= option is not specified
and the value of SINGULAR otherwise
Note: In many cases, a normalized matrix D-1AD-1
is decomposed and the singularity criteria are modified
correspondingly.
- If the matrix A is found singular in the first step,
a generalized inverse is computed. Depending on the G4=
option, either a generalized inverse satisfying all four
Moore-Penrose conditions is computed or a generalized
inverse satisfying only two Moore-Penrose conditions in
general. If the number of parameters n of the
application is less than or equal to G4=i, a G4 inverse
is computed; otherwise only a G2 inverse is computed.
The G4 inverse is computed by (the computationally very
expensive but numerically stable) eigenvalue decomposition,
the G2 inverse is computed by Gauss transformation.
The G4 inverse is computed using the eigenvalue
decomposition
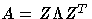 , where Z is the
orthogonal matrix of eigenvectors and
, where Z is the
orthogonal matrix of eigenvectors and 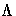 is
the diagonal matrix of eigenvalues,
is
the diagonal matrix of eigenvalues,
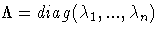 . If the PEIGVAL option is specified, the eigenvalues
. If the PEIGVAL option is specified, the eigenvalues
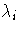 are displayed. The G4 inverse of A is set to
are displayed. The G4 inverse of A is set to
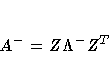
where the diagonal matrix
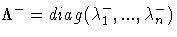 is defined using the COVSING= option
is defined using the COVSING= option
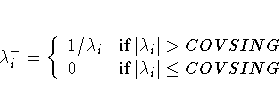
If the COVSING= option is not specified, the nr
smallest eigenvalues are set to zero, where nr is
the number of rank deficiencies found in the first step.
For optimization techniques that do not use second-order
derivatives, the covariance matrix is usually computed
using finite difference approximations of the derivatives.
By specifying TECH= NONE, any of the covariance matrices
can be computed using analytical derivatives. The covariance
matrix specified by the COV= option can be displayed
(using the PCOV option)
and is written to the OUTEST= or OUTVAR= data set.
Copyright © 1999 by SAS Institute Inc., Cary, NC, USA. All rights reserved.
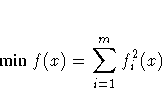
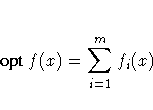
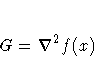
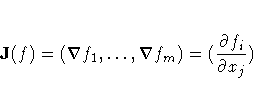
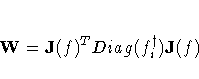
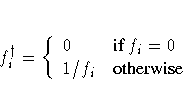
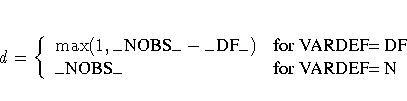
![]() depends on the specifications of the SIGSQ=
options and on the value of d:
depends on the specifications of the SIGSQ=
options and on the value of d:
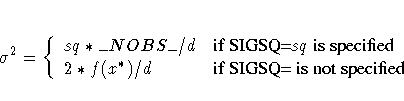
![]() is defined by
is defined by